When you are working with human, mouse, or rat targets─IDT has already designed qPCR sequences across these transcriptomes. We use an in silico design engine that screens your sequences for the ideal design. Select from this collection of sequences using the PrimeTime™ Predesigned qPCR Sequence Database.
When IDT predesigned human, mouse, or rat sequences are not suitable for your project─use the PrimerQuest™ Tool. This tool helps you design your custom assay by generating a list of ideal designs options using your nucleic acid sequences. This tool is powerful because its design engine incorporates calculations from thermodynamics research, and it offers you the option to customize approximately 45 different parameters, ranging from primer, probe, or amplicon criteria to target regions.
Primer designing tools
PrimeTime Predesigned qPCR Sequence Database Select primers/probes for qPCR (human, mouse, rat transcriptome).
- Predesigned sequences will result in 90% efficiency or better, or IDT will replace it with an alternative design, free of charge
- Sequence selection incorporates bioinformatic calculations that manage factors such as:
- Cross-react searches to avoid off-target amplification
- Recognition of splice variants
- Secondary structure predictions
- Orders include primers and probe sequences
PrimerQuest Tool Design primers or assays for PCR, qPCR, or sequencing (any species).
- Software allows customization of ~45 parameters, which can produce qPCR assay designs:
- With specific primer, probe, or amplicon criteria
- Across a specified location
- Around a fixed primer or probe location
- Design algorithm includes multiple checks to reduce primer-dimer formation
- Software provides flexible sequence entry and batch entries (up to 50 sequences)
The PrimerQuest Tool in 4 steps
The PrimerQuest Tool is the program of choice for designing qPCR primers/probes, sequencing oligonucleotides, and custom primers. Here is how to select the best PCR/qPCR assays for your experiments.
Step 1: Enter sequence(s)
The PrimerQuest Tool can perform batch analyses with up to 50 sequences that are each longer than 80 bases. You have 3 options for submitting your target sequence(s):
- Enter manually using FASTA format
- Download using Genbank or Accession ID
- Upload in an Excel file
Step 2: Choose your design
The PrimerQuest Tool offers 4 design options that are based on algorithms specific for common experimental setups (Figure 1). By default, your results return the 5 best primer or assay designs.
- PCR (2 primers)
- qPCR (2 primers + probe; for use in 5′ nuclease assays)
- qPCR (2 primers; for use with intercalating dyes)
- Custom (see Step 3 for additional information)

Step 3 (Optional): Customize your design parameters
Selecting Show Custom Design Parameters allows you to adjust reaction conditions (e.g., primer, Na+, and Mg2+ concentrations), force use of a particular primer or probe sequence, and/or specify design regions. Alternatively, if you originally chose PCR or qPCR designs, you may click on Customize Assay Design from your Results page to adjust design parameters (Figure 2). This step provides an overview of how to customize your primer designs:
1. Select your basic Design Parameters (Figure 2).
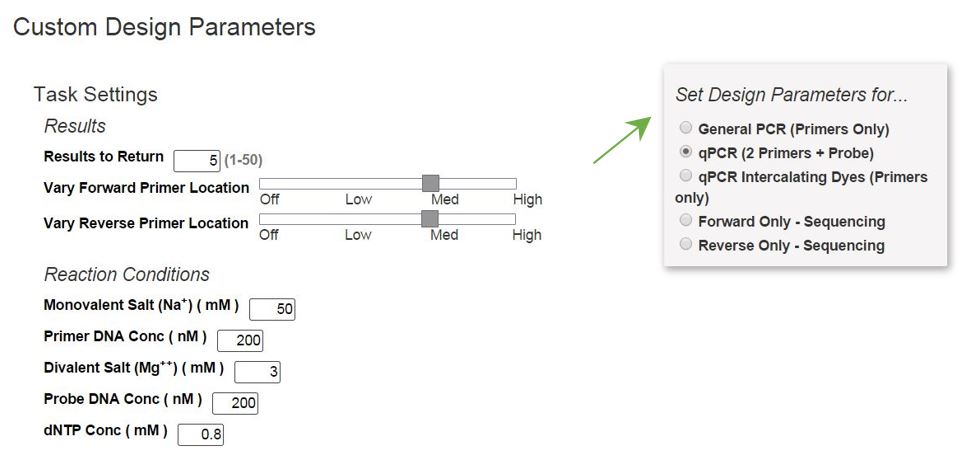
2. Customize at least 1 criterion (Table 1) used to design your primers and probe. Hints: Click on any criterion name to open a window with a definition of the criterion. If needed, select a Design Parameter radio button again to reset all the criteria back to default settings.
Table 1 contains descriptions of commonly customized parameters. Some criteria will only be visible for relevant Design Parameters. For example, probe parameters are only visible when designing oligos for qPCR with 2 primers and a probe.
| Criteria | Description |
|---|---|
| Results—results to return | The number of assay designs to view after clicking the Get Assays button (default: 5) Note: You cannot vary this parameter when doing a batch analysis. |
| Reaction conditions—divalent salt (Mg2+) (mM) | The millimolar concentration of divalent salt cations (usually MgCl2) in the reaction (range: 0–600 mM) |
| Primer criteria—primer Tm (°C) | Minimum, optimum, and maximum melting temperature for a primer oligo (range 45–75°C) |
| Primer criteria—primer GC (%) | Minimum, optimum, and maximum percentage of G and C bases in the primer (range: 20‑80%) |
| Amplicon criteria—amplicon size (bp) | The minimum, optimum, and maximum size for the desired amplicon |
3. Click the Get Assays button at the bottom of the web page.
4. If nothing is found, click on Adjust Parameters to modify criteria based on the information on the Results page, and then click the Get Assays button again.
Critical primer design parameters
There are ~45 customizable design parameters in the PrimerQuest Tool. Additional, fixed parameters exist to ensure the best primer or assay designs are generated. Here are short descriptions of a few important primer design parameters that are not variable:
- Poly-base runs are restricted to 3 consecutive, repeat bases or less to avoid polymerase slippage during primer extension.
- The difference in the Tm of the forward and reverse primers is always ≤3°C for reaction efficiencies.
- Probes cannot have a G base at the 5′ end because G bases partially quench most green/yellow dyes.
Step 4: Select and order assays
A schematic of your sequence is near the top of each Results page with the location of the assays (i.e., amplicons) depicted by green bars (Figure 3). The assay sets are listed below the schematic with the best assays at the top of the list. The list also includes important details about the primers or primers/probe, which can be downloaded as an Excel file if you are logged into your IDT account. Clicking on View Assay Details provides your submitted sequence along with highlights for the location of the primers or primers/probe.
If you are doing a batch analysis, you will find a dropdown menu to view the top 5 assays for each sequence (Figure 3).
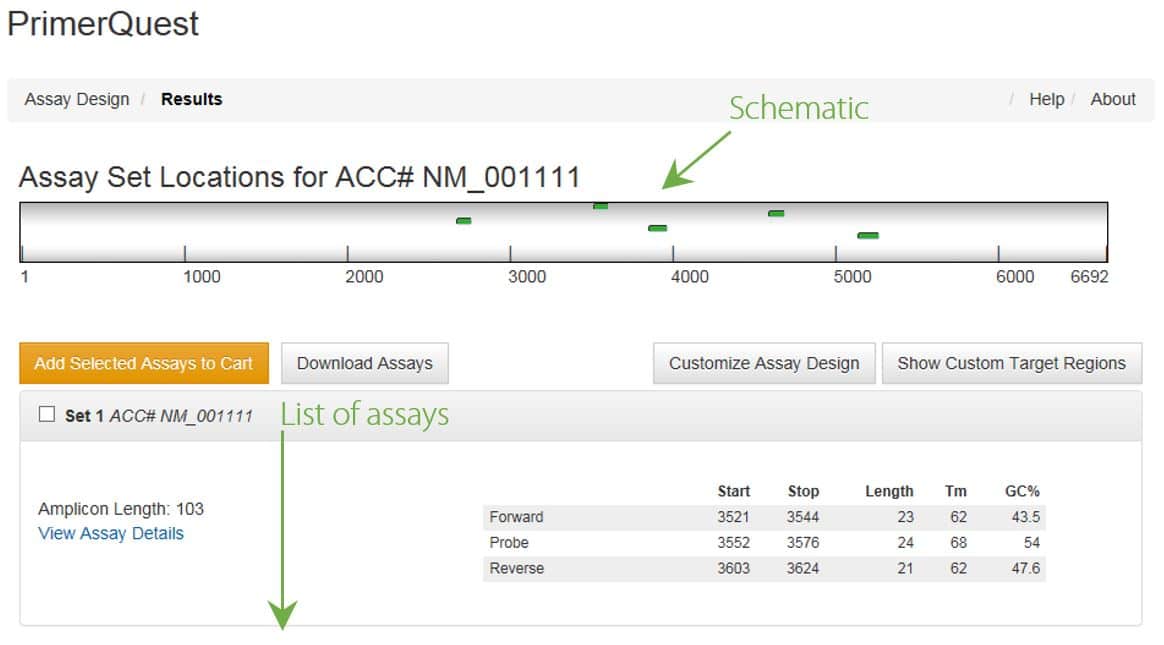
Use the following steps to select your assays:
- (Recommended) Test primer sequences for cross-reactivity with other known sequences using NCBI BLAST (Basic Local Alignment Search Tool), as predesigned sequences have not been previously checked using BLAST
- Use one of the following methods to select assays to add to your cart:
- Click on the box(es) next to the set number(s) that you wish to order.
- Click on the green assay bar(s) in the schematic.
- For batch analyses, click on the Select Top Result for All Sets box above the schematic.
- Click the Add Selected Assays to Cart button.
- Choose options (e.g., synthesis scale, purification options, fluorescent dyes, and quenchers) as needed, and follow screens to checkout.
More complex PCR/qPCR primer designs?
The PrimerQuest Tool helps to enable the design of basic and highly customized primers and probes for PCR and qPCR. Now that you understand the fundamentals of using this design tool, look for future articles from IDT about experiments requiring more complex assay design considerations, such as splice-specific, multiplexing, SNPs, and CNV designs.
If you need additional assistance with designing primers, contact us. Additionally, read our decoded article, How to Use BLAST to Locate PCR Primers.
For research use only. Not for use in diagnostic procedures. Unless otherwise agreed to in writing, IDT does not intend these products to be used in clinical applications and does not warrant their fitness or suitability for any clinical diagnostic use. Purchaser is solely responsible for all decisions regarding the use of these products and any associated regulatory or legal obligations. RUO23-1662_001