DNA methylation is an epigenetic mechanism that can regulate gene expression through several cell divisions, and even be heritable, without altering the underlying nucleotide sequence (see the sidebar, DNA methylation in vertebrates)[1]. A small but significant proportion of all CpG islands become methylated during development, and when this happens the associated promoter is stably silent. Thus, methylation provides a means for non-genetic factors to influence gene behavior. Environment, diet, stress, and aging are all factors that can cause changes in DNA methylation patterns. Some of these changes in epigenetic patterns are necessary, e.g., those involved in normal cellular differentiation; but they can also be detrimental, such as those that occur in many forms of cancer [1].
DNA methylation: From the whole genome to specific gene regions
Zymo Research Corporation, The Epigenetics Company (www.zymoresearch.com), provides kits and reagents for studying DNA methylation and hydroxymethylation patterns. Distinct DNA methylation patterns in the promoter regions of those genes may lead to changes in their expression. These analyses can be informative for classifying pairs of samples with distinct gene expression patterns for a certain set of genes, e.g., tumor suppressors in tumor and adjacent normal tissue.
The company introduced the OneStep qMethyl™ DNA Methylation Quantification Kit as a simple way to examine region-specific DNA methylation using methylation-sensitive restriction enzymes (MSREs) and probe-based qPCR. The method does not measure the individual methylation status of each CpG (as in bisulfite sequencing), but rather provides an average DNA percent methylation for that particular region covered by the amplicon.
This information can be sufficiently instructive for researchers looking candidate biomarkers. Furthermore, the MSRE and real-time PCR method is a quick and inexpensive means for researchers with a background in real-time PCR to analyze DNA methylation differences in a specific subset of genes or CpG islands within specific promoters; thus, avoiding whole genome sequencing.
MSRE workflow
Genomic dsDNA is digested by restriction enzymes that cleave unmethylated cytosines in the DNA. Where cytosines contained in specific restriction sites are methylated, the sequence remains intact. Real-time PCR is then used to amplify regions containing these CpG sites, and the sizes of the amplicon fragments recovered determine the average methylation state.
Intact, methylated regions show high levels of amplification, whereas amplicons containing unmethylated cytosines at those restriction sites show late amplification, essentially as noise due to small amounts of nonspecific primer binding, as with the No Template Control sample. Figure 1 provides a more detailed description of the procedure and the formula used to calculate percent methylation.
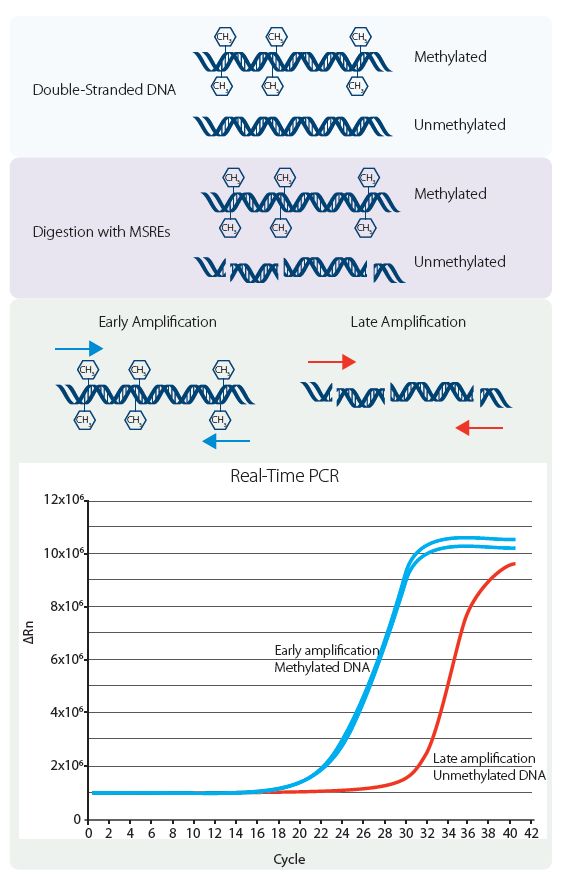
Zymo Research offers both a fluorescence-based singleplex system (the SYTO9) and a version of the procedure for use with a probe for multiplexing. A major advantage of using the probe-based system is that unmethylated amplicons are amplified much later, and this provides better discrimination between methylated and unmethylated fragments than traditional methylation sequencing.
Primer design is also a crucial part of this workflow. IDT offers a free online tool for PCR and qPCR primer design that provides flexible sequence entry and batch entries up to 50 sequences. To find out more check out the PrimerQuest Tool web page. You can also design Custom PrimeTime qPCR Assays for this application.
Obtaining truly unmethylated standards
Use of genomic methylated and non-methylated standards, available from Zymo Research, provides a method to quantify methylation on a global level. For specific gene regions of any species, the company uses IDT gBlocks Gene Fragments to obtain double-stranded sequences that are completely unmethylated, and which can then be custom methylated.
Keeping DNA methylation simple
Zymo Research keeps their approach for evaluating methylation simple. The OneStep qMethyl Kits are a great way for researchers to explore differentially methylated regions in the genome as a first step, before moving to targeted sequencing or another method that requires bisulfite conversion of the DNA.
References
- Moore LD, Le T, Fan G. DNA methylation and its basic function. Neuropsychopharmacology. 2013;38(1):23-38.
- Angeloni A, Bogdanovic O. Sequence determinants, function, and evolution of CpG islands. Biochem Soc Trans. 2021;49(3):1109-1119.
For research use only. Not for use in diagnostic procedures. Unless otherwise agreed to in writing, IDT does not intend these products to be used in clinical applications and does not warrant their fitness or suitability for any clinical diagnostic use. Purchaser is solely responsible for all decisions regarding the use of these products and any associated regulatory or legal obligations. Doc ID: RUO23-2012_001
























