Adapters for next generation sequencing
High quality adapters that meet the ever-expanding needs of NGS researchers
Adapters are a key component of the next generation sequencing (NGS) workflow. Whether your project requires a basic adapter or a sophisticated design for higher accuracy, we have the products and expertise to deliver the right solution.
Regardless of the NGS instrument or application, we have been serving the needs of NGS scientists for over 10 years. We are recognized as the leader in adapter synthesis based on our experience in custom oligo manufacturing and our commitment to quality. Our NGS customers and partners benefit from:
- A comprehensive adapter offering
- Innovative designs
- Trusted customer support
- Complete customizability
Our leadership in NGS adapter synthesis made us the clear choice when Illumina sought a partner to develop the next generation of index adapters to improve sample multiplexing. These adapters contain Illumina’s unique dual indexes (UDIs), which mitigate sample misassignment due to index hopping. We manufacture UDI adapters for Illumina and are also licensed to sell your own custom designs containing these UDI sequences.
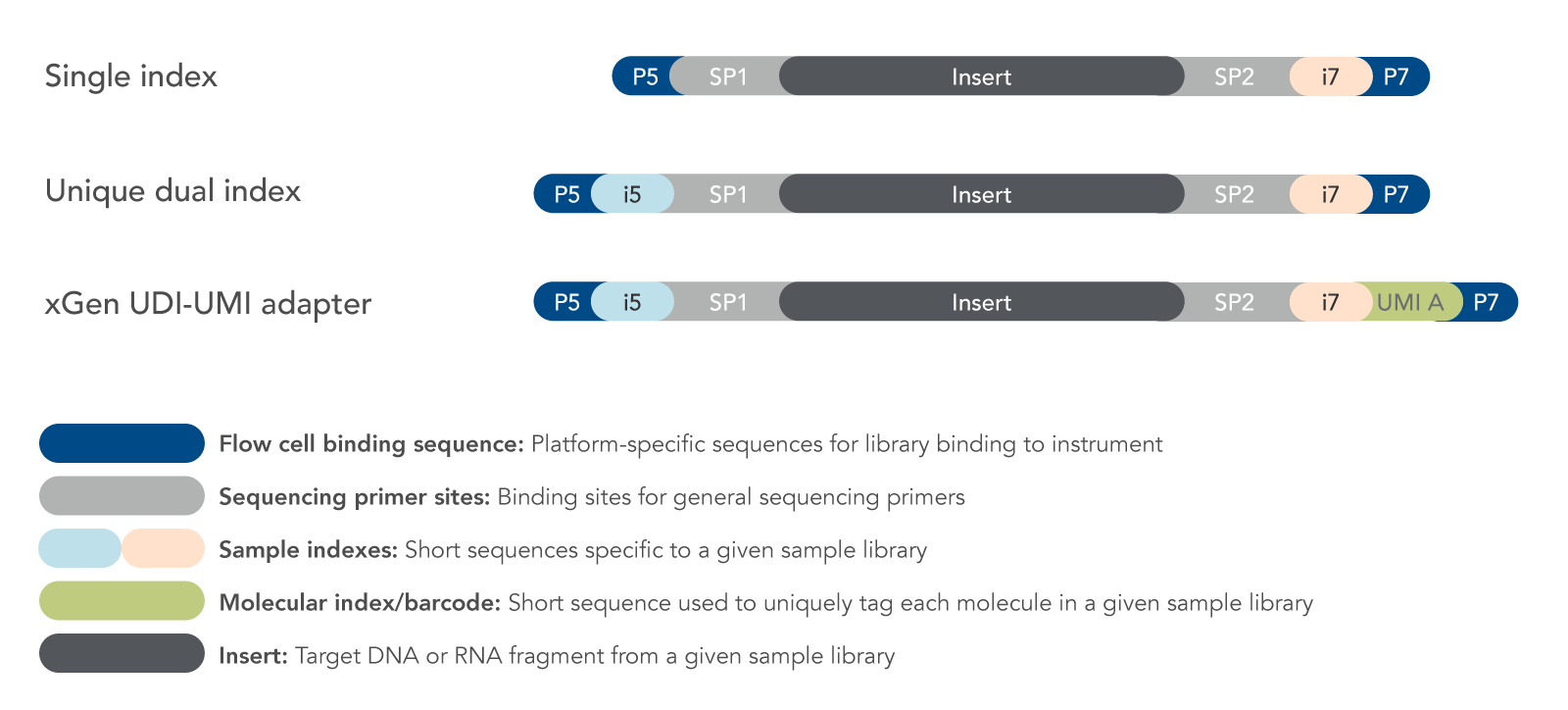
Figure 1. Examples of adapter designs. A variety of NGS adapter designs are available. When selecting or designing adapters, consideration must be given to the intended application, multiplexing needs, accuracy requirements, and analysis methods.
Considerations when selecting an adapter design
IDT provides xGen™ UDI-UMI Adapters, xGen Stubby Adapter and Unique Dual Index Primers, and a Custom Adapter Configurator tool that guides you through design of Custom NGS Adapters. Below, we describe important design considerations you will want to take into account.
Sequences for specific NGS platforms: During library preparation, adapters are attached (by ligation, PCR, or tagmentation) to the fragments of each sample library. Adapters include platform-specific sequences for fragment recognition by the sequencing instrument: for example, the P5 and P7 sequences (Figure 1) enable library fragments to bind to the flow cells of Illumina platforms. Each NGS instrument provider uses a specific set of sequences for this purpose. IDT manufactures adapters for all major NGS platforms.
Sample indexing: Sample indexes (or indices) enable multiple samples to be sequenced together (i.e., multiplexed) on the same instrument flow cell or chip. Each sample index, typically 8–10 bases, is specific to a given sample library and is used for de-multiplexing during data analysis to assign individual sequence reads to the correct sample. Predesigned adapters from IDT are base-balanced to prevent GC bias and color-balanced for 2 and 4-color Illumina instruments. Adapters may contain single or dual sample indexes depending on the number of libraries combined and the level of accuracy desired. Illumina recommends using unique dual indexes (UDIs) as a method to mitigate errors introduced by index-hopping. UDIs are particularly important when using instruments with patterned flow cells, such as the NovaSeq system.
IDT provides several indexing options with its Custom NGS Adapters. These include both IDT and Illumina 8 and 10 base index series, as well as the ability to upload your own index files.
Molecular barcoding: Unique molecular identifiers (UMIs) provide the highest levels of error correction and accuracy. UMIs are short sequences, often with degenerate bases, that incorporate a unique barcode onto each molecule within a
given sample library. UMIs enable precise quantification, since PCR duplicates will share the same insert and UMI tag. UMI deduplication is useful for RNA-seq gene expression analysis and other quantitative sequencing methods. UMIs have also been
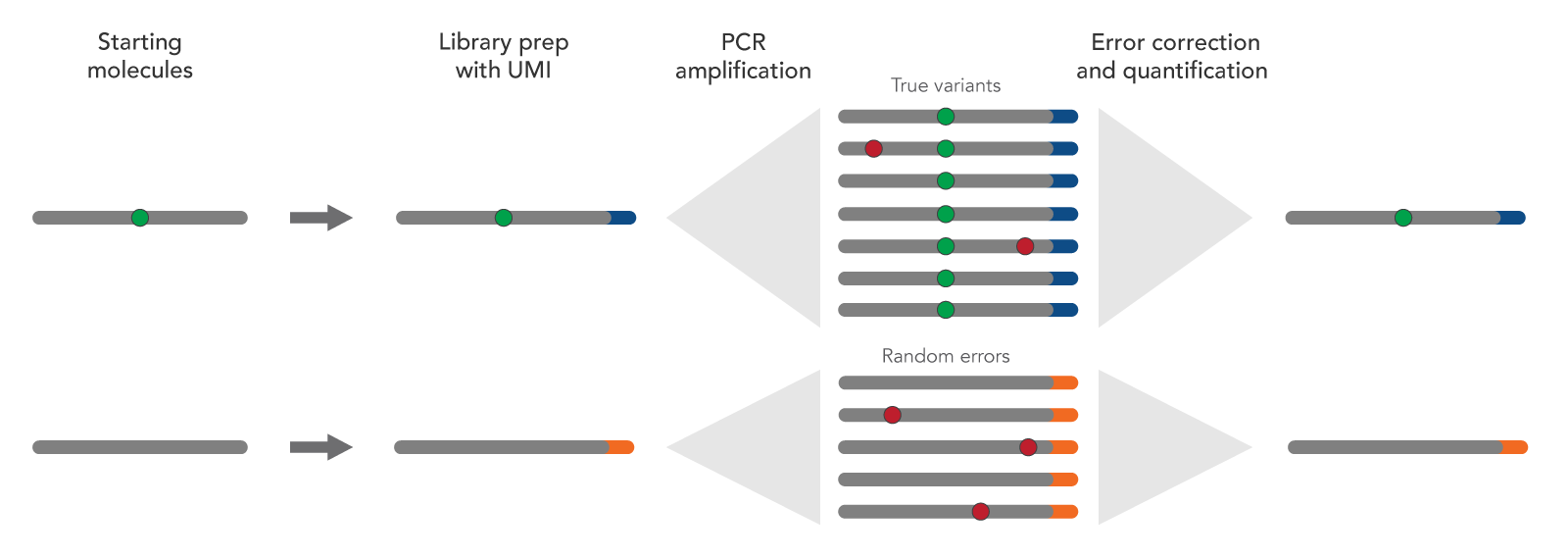
shown to reduce the rate of false-positive variant calls and increase sensitivity of variant detection. By incorporating individual barcodes on each original DNA fragment, variant alleles present in the original sample (true variants) can be distinguished
from errors introduced during library preparation, target enrichment, or sequencing (Figure 2). Any identified errors can be removed by bioinformatics methods before final data analysis. Adapters that contain UMIs, such as the xGen UDI-UMI Adapters, are available with a UDI design for detection of low-frequency variants. IDT Custom NGS Adapters can also be configured with UMIs.
Adapter manufacturing: Stringent manufacturing methods are critical for producing high quality NGS adapters. Substandard manufacturing practices can lead to low purity adapters or adapter cross-contamination, either of which will negatively impact sequencing results. The IDT proprietary TruGrade™ process uses state-of-the-art synthesis and purification methods, designed specifically for NGS adapters. IDT also offers GMP grade adapter manufacturing for clinical research applications.
Choose IDT adapter products based on your experimental needs. The Custom NGS Adapters provide a step-by-step tool to help you configure adapters best suited for your research.
| Specifications | xGen UDI-UMI Adapters | xGen Stubby Adapter and UDI Primers | Custom NGS adapters |
|---|---|---|---|
| Configuration | UDI-UMI | UDI | single index, combinatorial dual indexing, UDI, or UDI-UMI |
| Sample index length | 8 base, unique sample indexes | 8 base, unique sample indexes | Configurable |
| Concentration | 15 µM | 15 µM (Stubby Adapter), | 15 µM (Adapter), 20 µM total (10 µM each Primer) |
| Shipping container | 96-well plates | Tubes (Stubby Adapter), | Tubes, Matrix® plates†, or single-use plates |
| Reactions | 1 reaction per adapter | 1 reaction per primer pair | Varies |
| Methyl-Seq compatible | Available | Methylated for bisulfite sequencing* | Available |
| Illumina platform compatibility | 2- and 4-color platforms | 2- and 4-color platforms | Designs for any application |
* For bisulfite sequencing, see our FAQ for additional information.
† Matrix is a trademark of Thermo Fisher
Scientific.
IDT is an authorized re-seller of Illumina’s unique dual indexes for custom adapter designs.
For more information, or to order custom adapters, please contact our Technical and Customer Support.
Product pages
xGen UDI-UMI Adapters »
Single-use, TA-ligation-ready, full-length adapters
xGen Stubby Adapter and UDI Primers »
Single-use, TA-ligation-ready, stubby Y adapter and indexing primers
Custom NGS Adapters »
Adapter and indexing primer selections for DNA library prep


 Processing
Processing