Frequently asked questions
Our team has assembled a list of frequently asked questions to help you find answers quickly. Filter using one or more categories to focus on specific topics, or use the search bar to perform a text search.
Search all FAQs:
How can I ensure I am designing my crRNA protospacer sequence in the correct orientation and to the appropriate strand?
Because the crRNA recognizes and binds 19–20 nt on the genomic DNA strand opposite to the NGG sequence of the PAM site, your crRNA protospacer sequence should include the 19–20 nt upstream of the PAM site, in the forward orientation. See Figure 2 on the Overview tab of the Alt-R™ CRISPR-Cas9 System product page for an example. Make sure you verify the correct strand orientation of the protospacer sequence, and do not include the PAM site. Common incorrect design examples are shown in red in the figure below.
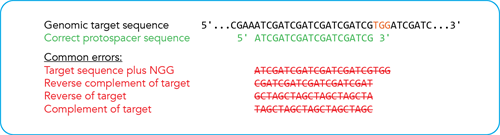
How to enter your crRNA target sequence. Because the crRNA recognizes and binds 20 bases on the DNA strand opposite from the NGG sequence of the PAM site, order your crRNA by entering the 20 bases upstream of the PAM site, in the forward orientation as shown. If you are pasting your CRISPR target site from an online design tool, make sure you verify the correct strand orientation. Do not include the PAM site. Common incorrect design examples are shown in red.