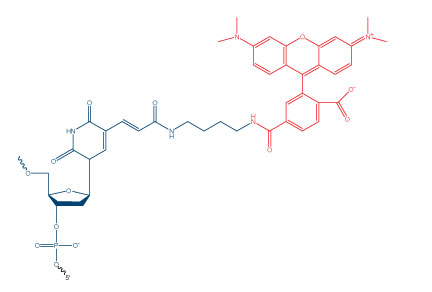
Many researchers are unaware that certain modifications may require the presence or addition of a specific nucleotide in their oligonucleotide sequence. These modifications come in many forms, including fluorophores, spacers, and attachment chemistry/linker modifications. The most common nucleotide that is introduced with these modifications is thymidine. Some of these modifications have “dT” included in their name, such as 3’ Biotin dT or Internal Fluorescein dT, while others are more difficult to identify. For example, Internal TAMRA NHS Ester (Figure 1) does not include dT in its name but is attached after oligonucleotide synthesis via an amino dT.
All internal NHS Esters and click-enabled modifications are attached through a thymidine residue after oligo synthesis. While most 3’ and 5’ modifications, including NHS Esters and click modifications, do not add an additional base to the oligo, 3’TAMRA™ in phosphoramidite form is one exception to this rule, as it is built from a dT residue.
Including a modification code (e.g., an internal TAMRA attachment) in your sequence during the ordering process will add a dT residue at that position. Thus, the modification code /i6-TAMN/ within the sequence TCGA/i6-TAMN/CGTA will incorporate a “T” nucleotide at that position. The final sequence will be: TCGA “T” CGTA, with TAMRA attached to the additional dT residue. It is therefore important for you to account for the necessary dT base in your sequence prior to ordering, i.e., when assessing binding to a target sequence and in any calculations, such as melting temperature (Tm).
Review a list of the common modifications IDT can add to oligonucleotides here. Not finding a modification you need on the IDT website? No worries. IDT will consider any modification you need, just contact us.
For research use only. Not for use in diagnostic procedures. Unless otherwise agreed to in writing, IDT does not intend these products to be used in clinical applications and does not warrant their fitness or suitability for any clinical diagnostic use. Purchaser is solely responsible for all decisions regarding the use of these products and any associated regulatory or legal obligations. Doc ID: RUO23-1934_001

