Introduction
CRISPR gene editing continues to be an exciting and powerful tool for scientific research. Though introducing gene disruptions by CRISPR has become a simple endeavor, inserting precise sequences via homology-directed repair or “knock-in” can still be a challenge. Even with tool-aided design of guide RNA and HDR templates, HDR rates can be frustratingly low, delaying the time it takes to attain the right genomic context for scientific studies, often stalling project progress.
Experiment
Here at IDT, we focused on addressing HDR inefficiency. Our research scientists previously conducted experiments on the mechanism of CRISPR HDR and developed a small molecule reagent, Alt-R HDR Enhancer, that effectively diverts the double strand break repair pathway, significantly increasing HDR efficiency in many common cell lines. Since launching this product in 2018, we have seen great achievement and had positive reviews from customers doing HDR in a variety of cell types. During that span of time, our scientists continued to research small molecules that could potentially outperform even our own product. We selected a large number of these small molecules for a screening experiment.
Screening of the selected small molecules was performed as follows. Ribonucleoprotein (RNP) complex (2 µM) targeting human HPRT1 and 3 µM Alt-R HDR Donor designed to insert an EcoRI restriction site were delivered together into Jurkat cells by electroporation using a 4D-Nucleofector™ System (Lonza). Immediately after electroporation, cells were cultured in media containing no addition, DMSO (vehicle control), or one of the following potential small-molecule HDR enhancers: 0.02 µM wortmannin, 5 µM SCR7, 5 µM L755507, 0.1 µM EPZ5676, 5 µM rucaparib, 1 µM pevonedistat, 0.1 µM brefeldin A, 30 µM Alt-R HDR Enhancer V1, 10 µM XL413, 2.5 µM NU7441, 10 µM trichostatin A, "CRISPY mix" (30 µM NU7026, 10 µM trichostatin A, 1 µM MLN4924), 0.1 µM romidepsin, 20 µM nedisertib (M3814), or 1 µM Alt-R HDR Enhancer V2. Sources and references for these small molecules are listed in Table 1.
Table 1. Source or reference used for molecules in our screening experiment.
| Small molecule name | Concentration (µM) | Reference/source |
|---|---|---|
| Wortmannin | 0.02 | Delacote et al. [1] |
| SCR7 | 5 | Maruyama et al. [2] |
| L755507 | 5 | Yu et al. [3] |
| EPZ5676 | 0.1 | Stein et al. [4] |
| Rucaparib | 5 | Iyer et al. [5] |
| Pevonedistat/MLN4924 | 1 | Riesenberg et al. [6] |
| Brefeldin A | 0.1 | Yu et al. [3] |
| Alt-R HDR Enhancer (V1) | 30 | IDT |
| XL413 | 10 | Wienert et al. [7] |
| NU7441 | 2.5 | Leahy et al. [8] |
| Trichostatin A | 10 | Riesenberg et al. [6] |
| "CRISPY mix" | 30 µM NU7026, 10 µM trichostatin A, 1 µM MLN4924 | Riesenberg et al. [6] |
| Romidepsin | 0.1 | Bertino et al. [9] |
| Nedisertib/M3814 | 20 | Fu et al. [10] |
| Alt-R HDR Enhancer V2 | 1 | IDT |
Genomic DNA was isolated 48 hr after transfection, and target regions were amplified by PCR and sequenced on a MiSeq® (Illumina). HDR efficiency was quantified using the rhAmpSeq CRISPR Analysis Tool. As shown in Figure 1, we identified one small molecule, Alt-R HDR Enhancer V2, that enhanced HDR efficiency more than any of the other small molecules in the screening experiment. The highest HDR was achieved when using the Alt-R HDR Enhancer V2 at 1 µM.
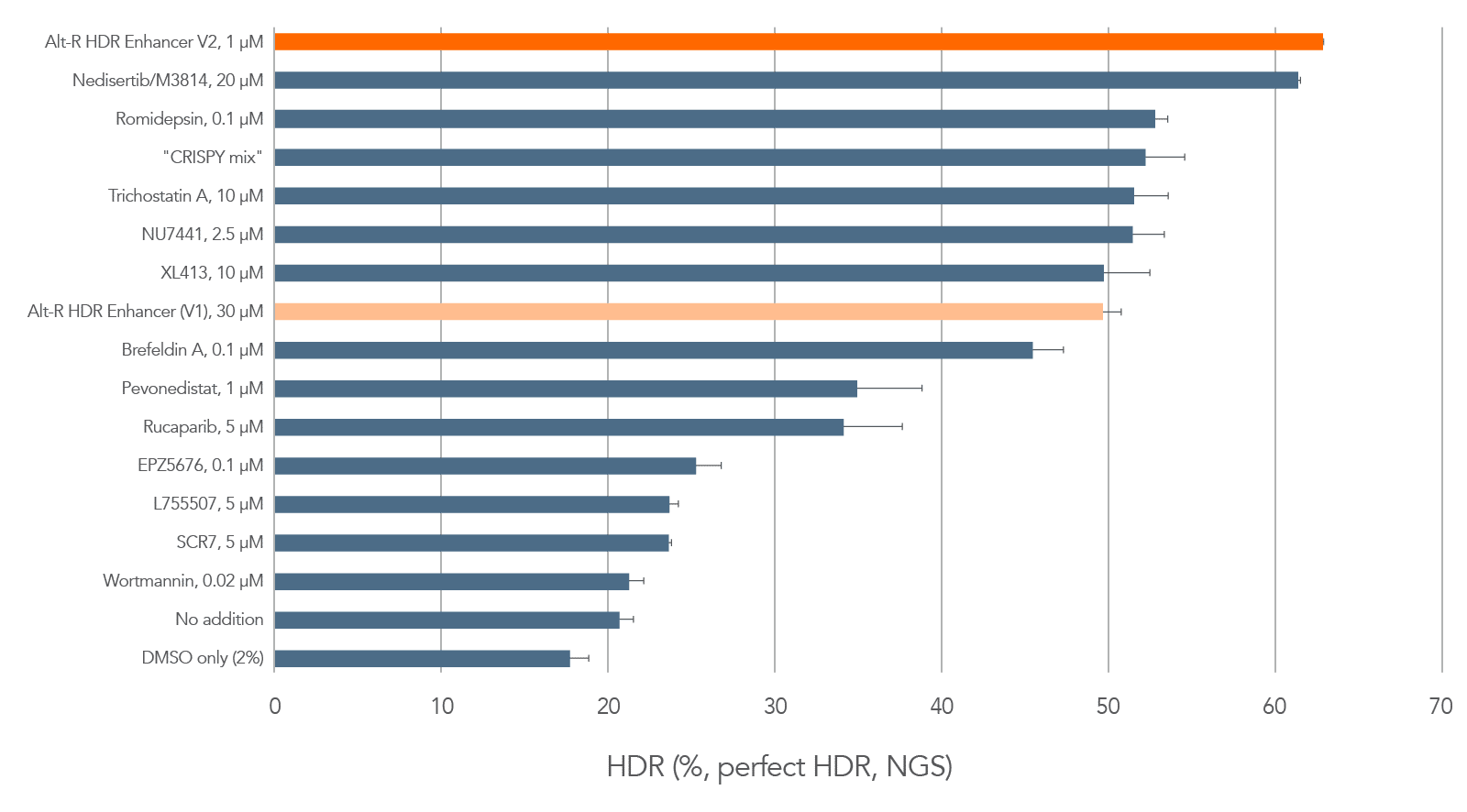
Alt-R HDR Enhancer V2 improved HDR efficiency in many commonly used human cell lines, including primary T cells [11], Jurkat, HEK-293, HeLa, U2OS, HAP1, K562, iPSCs, Hepa1-6, B16-F12, and H36.12j compared to untreated counterparts (data not shown). This enhancement of HDR efficiency was observed following both Cas9 and Cas12a Ultra genome editing. See the product page for more information.
Conclusion
The new Alt-R HDR Enhancer V2 is a small molecule that increases HDR efficiency in a broad range of cultured human cell lines compared to untreated cells. It can be used at a concentration as low as 1 µM, compared to the approximately 30 µM concentrations needed for the original version of Alt-R HDR Enhancer. Therefore, Alt-R HDR Enhancer V2 is expected to provide wide utility in cell culture HDR experiments.
References
- Delacote F, Han M, Stamato TD, et al. An xrcc4 defect or Wortmannin stimulates homologous recombination specifically induced by double-strand breaks in mammalian cells. Nucleic Acids Res. 2002;30(15):3454-3463.
- Maruyama T, Dougan SK, Truttmann MC, et al. Increasing the efficiency of precise genome editing with CRISPR-Cas9 by inhibition of nonhomologous end joining. Nat Biotechnol. 2015;33(5):538-542.
- Yu C, Liu Y, Ma T, et al. Small molecules enhance CRISPR genome editing in pluripotent stem cells. Cell Stem Cell. 2015;16(2):142-147.
- Stein EM, Garcia-Manero G, Rizzieri DA, et al. A Phase 1 Study of the DOT1L Inhibitor, Pinometostat (EPZ-5676), in Adults with Relapsed or Refractory Leukemia: Safety, Clinical Activity, Exposure and Target Inhibition. Blood. 2015;126(23):2547-2547.
- Iyer S, Suresh S, Guo D, et al. Precise therapeutic gene correction by a simple nuclease-induced double-stranded break. Nature. 2019;568(7753):561-565.
- Riesenberg S, Maricic T. Targeting repair pathways with small molecules increases precise genome editing in pluripotent stem cells. Nat Commun. 2018;9(1):2164.
- Wienert B, Nguyen DN, Guenther A, et al. Timed inhibition of CDC7 increases CRISPR-Cas9 mediated templated repair. Nat Commun. 2020;11(1):2109.
- Leahy JJ, Golding BT, Griffin RJ, et al. Identification of a highly potent and selective DNA-dependent protein kinase (DNA-PK) inhibitor (NU7441) by screening of chromenone libraries. Bioorg Med Chem Lett. 2004;14(24):6083-6087.
- Bertino EM, Otterson GA. Romidepsin: a novel histone deacetylase inhibitor for cancer. Expert Opin Investig Drugs. 2011;20(8):1151-1158.
- Fu YW, Dai XY, Wang WT, et al. Dynamics and competition of CRISPR-Cas9 ribonucleoproteins and AAV donor-mediated NHEJ, MMEJ and HDR editing. Nucleic Acids Res. 2021;49(2):969-985.
- Kath J, Du W, Thommandru B, et al. Fast, efficient and virus-free generation of TRAC-replaced CAR T cells. bioRxiv. 2021:2021.2002.2014.431017
























