xGen™ SARS-CoV-2 Hybridization Panel
Strain-agnostic enrichment of SARS-Cov-2 genome
The xGen SARS-CoV-2 Hyb Panel spans the complete viral genome and is not susceptible to known and novel mutations arising from diverse strains.
xGen NGS—made for SARS-CoV-2 research.
Ordering
- Identify known and novel mutations arising from diverse strains
- Available as a bundle with xGen™ Universal Blockers-TS Mix and xGen Hybridization and Wash Kit
Transform Your NGS Workflow with Automation
Looking to streamline your NGS workflows? Discover how automation can enhance efficiency and consistency in your lab with our NGS Automation solutions.
For the xGen Hybridization and Wash Kit, xGen Universal Blockers, xGen Library Amplification Primer Mix, and/or xGen Human Cot DNA, please visit the xGen Hybridization Capture Core Reagents page.
Product data
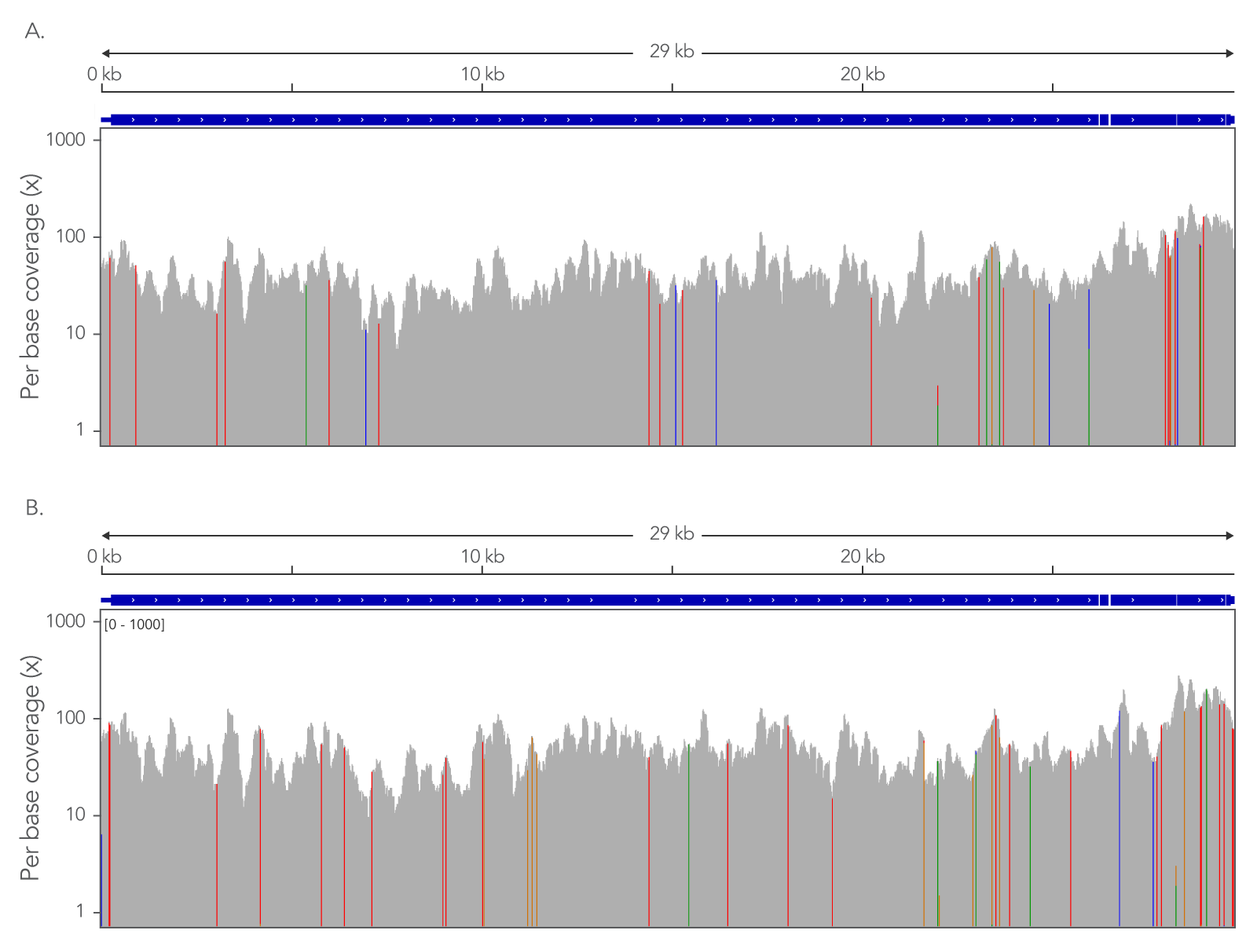
Even coverage across the complete viral genome
Figure 1. An example of the xGen™ SARS-CoV-2 Hyb Panel providing even coverage across the complete viral genome allowing strain identification with minimal read depth. (A) Coverage of a nasopharyngeal swap sample identified as belonging to strain Alpha (B.1.1.7). (B) Coverage of a nasopharyngeal swap sample identified as belonging to strain Delta (B.1.617.2). 5 µL of total RNA sample (previously shown to contain viral sequence via qPCR assay) were used to prepare NGS libraries (n=7) using the xGen™ Broad Range Library Prep Kit. Libraries were quantified and equal amounts were captured with the xGen™ Hybridization and Wash Kit. Samples were sequenced on an Illumina MiSeq™ and subsampled to 25000 total reads per sample. Data was aligned to the Refseq sequence (NC_045512) and viewed in the Integrated Genome Viewer (IGV). The IGV view color codes bases that do not match the canonical sequence. Strains were identified by submitting a consensus sequence for the sample to https://pangolin.cog-uk.io/