xGen™ NGS Adapters & Indexing Primers for Illumina® sequencing
High-quality stocked and customizable options
Adapters & indexing primers are a key component of NGS library preparation workflow. Whether your project requires a basic indexing solution or a more sophisticated design for higher accuracy, we have the products and expertise to deliver the right solution.
xGen NGS—made to differentiate.
Ordering
- IDT is the industry standard for both ligation-based and PCR-based indexing solutions
- A comprehensive offering of high-purity adapters and primers for Illumina sequencing
- Trusted customer support
- Complete customizability
Table 1. Recommended adapters and indexing primers for Illumina sequencing based on compatibility with library prep methods.
| Full-length adapters | Indexing primer options | |||||
|---|---|---|---|---|---|---|
| Indexing strategy | UDI + UMI | UDI | CDI | Normalase™-UDI* | Normalase-CDI* | |
| xGen Broad-Range RNA Library Prep Kit | ✓ | ✓ | ✓ | ✓ | ||
| xGen RNA Library Prep Kit | ✓ | ✓ | ✓ | ✓ | ||
| xGen DNA Library Prep (MC or EZ) Kit | ✓ | ✓ | ✓ | ✓ | ||
| xGen DNA Library Prep (MC or EZ) UNI† Kit | ✓ | |||||
| xGen ssDNA & Low-Input DNA Library Prep Kit | ✓ | ✓ | ✓ | ✓ | ||
| xGen Methyl-Seq Library Prep Kit | ✓ | ✓ | ✓ | ✓ | ||
| xGen cfDNA &FFPE DNA Library Prep Kit | ✓ | ✓ | ||||
* Index primers are also used for Pre-Hyb Normalase Workflow
† Note: the UNI kit supports indexing by ligation, so either full-length (UDI-UMI) or stubby adapter + UDI primers can be used. All other library prep kits only support indexing by PCR
For information on the xGen Normalase Module, click here.
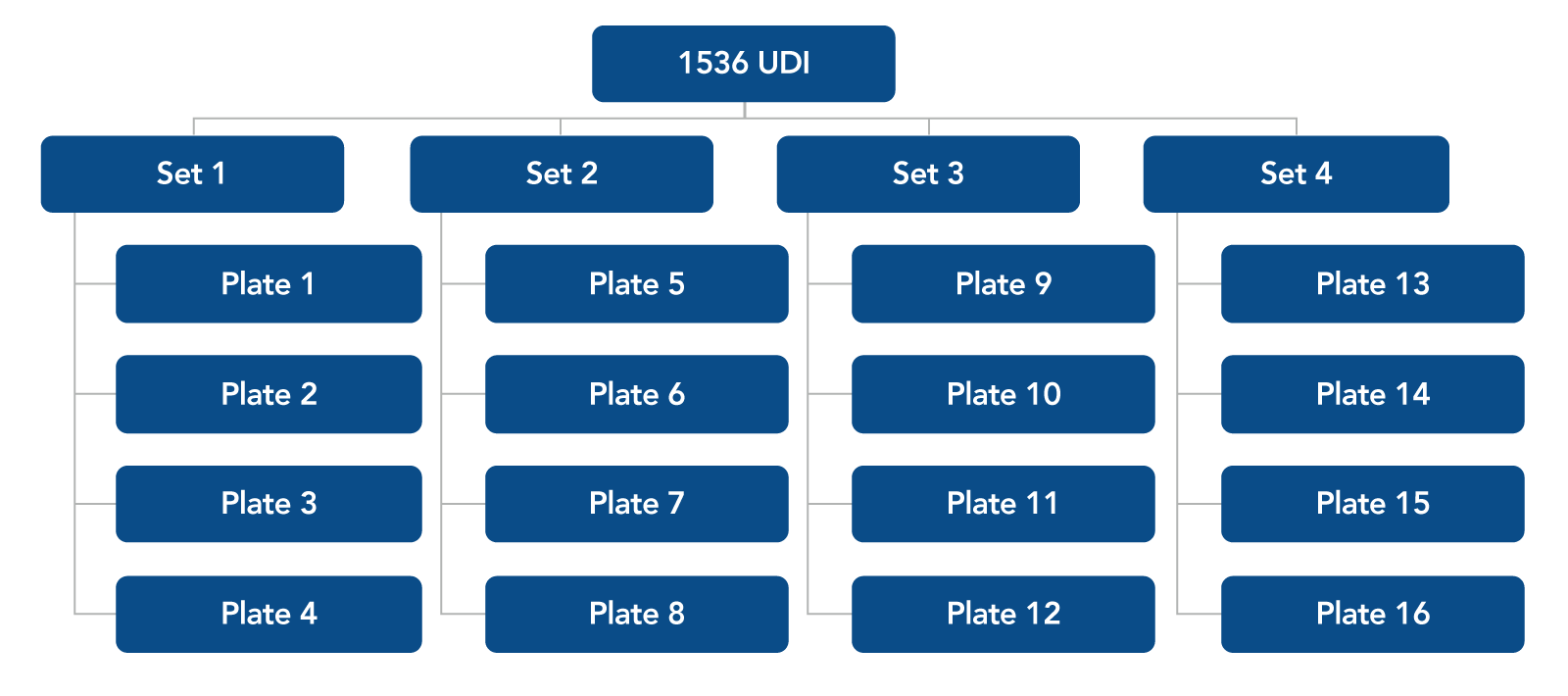
Figure 1. Plate combinations for 1536 UDI primer pairs. Each 96-well plate is sold in a set of four plates. For 1536 UDI primers, order Set 1, 2, 3, and 4.
For plate layouts of each of the 16 UDI plates offered see the xGen™ Normalase™ Indexing Primers quick reference guide.
Custom Adapter Configurator tool
Use the Custom Adapter Configurator tool to design indexing solutions for use with Illumina sequencing platforms, to take your research to the next level. These configurable options include library prep method, configuration, attachment style, indexing strategy and number of indexes, amount, delivery format, and whether to include modifications or TruGrade™ processing. The tool allows you to select adapters for immediate purchase through our online cart or request customer support for assistance if you have more complex requirements for your design.
Note: We suggest to avoid using custom adapters (such as those through the Custom Adapter Configurator Tool) and off-the-shelf, stocked adapters together in the same project. Please contact us for further assistance.
TruSeq™–Compatible Full-length Adapters
Each sequence design includes duplexed adapters, delivered in Duplex Buffer (IDT), and available in tubes or plates.
* Price per number of adapters selected. Addition of methylation and/or TruGrade processing will incur an additional fee.
TruSeq™–Compatible Stubby Adapter and Indexing Primers
Each sequence design includes duplexed adapters designed to be compatible with ligation-based library prep, delivered in Duplex Buffer (IDT) and provided in tubes, as well as a primer mix delivered in IDTE (1X TE), pH 8, available in plates.
* Price per number of adapter-primer sets selected. Addition of methylation and/or TruGrade processing will incur an additional fee.
Nextera™–Compatible Indexing Primers
Each sequence design includes a primer mix designed to be compatible with transposase-based library prep, delivered in IDTE (1X TE), pH 8, available in plates.
*Price per number of primer sets selected. Addition of TruGrade processing will incur an additional fee.
TruSeq™–Compatible Indexing Primers
Each sequence design includes a primer mix designed to be compatible with ligation-based library prep, delivered in IDTE (1X TE), pH 8, available in plates.
* Price per number of primer sets selected. Addition of TruGrade processing will incur an additional fee.
Additional fees for methylated bases
Additional fees for TruGrade processing
If ordering >384 sequences, please use this request form.
TruSeq® and Nextera® are registered trademarks of Illumina, Inc., used with permission. All rights reserved.
Product details
Considerations when selecting an adapter and primer design
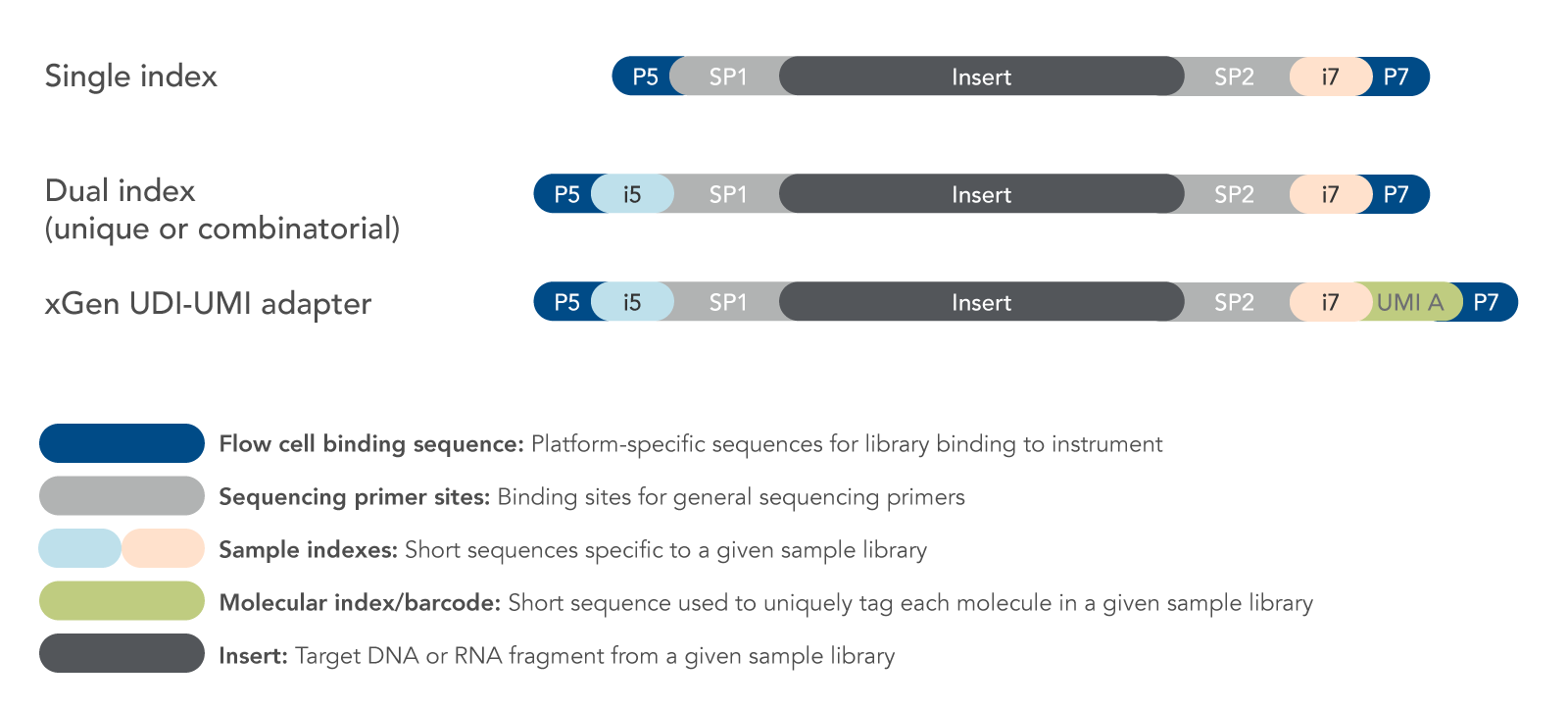
IDT provides extensive solutions for stocked indexing solutions. Adapters are available with single index, unique dual index (UDI), and unique molecular identifiers (UMI) found in the IDT xGen UDI-UMI indexes (Figure 2). In addition, the adapters can be designed to include:
- Index lengths of 8 nt or 10 nt
- Combinatorial or unique dual indexes (CDI or UDI)
- Multiplex capacity up to 1536 unique indexes
- Indexing primers designed for performing enzymatic library normalization using our proprietary Normalase technology.
In addition to the stocked products, IDT provides the ability for even greater flexibility and control, by using the Custom Adapter Configurator tool to guide you through designing your own Custom NGS Adapters.
Figure 2. Examples of adapter designs. A variety of NGS adapter designs are available from IDT. When selecting or designing adapters, consideration must be given to the intended application, multiplexing needs, specification requirements, and analysis methods.
Sample indexing
Sample indexes (or indices) enable multiple samples to be sequenced together (i.e., multiplexed) on the same instrument flow cell or chip. Each sample index, typically 8–10 bases, is specific to a given sample library and is used for de-multiplexing during data analysis to assign individual sequence reads to the correct sample. Predesigned indexes from IDT are balanced to prevent GC bias and color-balanced for 2- and 4-color Illumina instruments. Adapters and primers may contain single or dual sample indexes depending on the number of libraries combined and the level of specification desired. Illumina recommends using unique dual indexes (UDIs) as a method to mitigate errors introduced by index-hopping. UDIs are particularly important when using instruments with patterned flow cells, such as the NovaSeq® system from Illumina.
IDT provides several indexing options with its Custom NGS Adapters. These include both IDT and Illumina 8- and 10-base index series, as well as the ability to upload your own index files.
Molecular barcoding
Unique molecular identifiers (UMIs) provide reliable error correction. UMIs are short sequences that incorporate a unique barcode onto each molecule within a given sample library.
- UMIs are used to identify PCR duplicates. PCR duplicates created during library amplification steps will share the same insert and UMI tag.
- UMIs have also been shown to reduce the rate of false-positive variant calls and increase variant detection. By incorporating individual barcodes on each original DNA fragment, variant alleles present in the original sample (true variants) can be distinguished from errors introduced during library preparation, target enrichment, or sequencing. Any identified errors can be removed by bioinformatics methods before final data analysis.
Adapters that contain UMIs, such as the xGen UDI-UMI Adapters, are available with a UDI design for identification of low-frequency variants. IDT Custom NGS Adapters can also be configured with UMIs.
Adapter manufacturing
Stringent manufacturing methods are critical for producing high-quality NGS adapters. Substandard manufacturing practices can lead to low-purity adapters or adapter cross-contamination, either of which will negatively impact sequencing results. The IDT proprietary TruGrade™ process uses state-of-the-art synthesis and purification methods, designed specifically for NGS adapters. IDT also offers GMP ISO 13485 grade adapter manufacturing for research applications.
In addition to stringent manufacturing methods, IDT also performs functional NGS testing on each new batch of stocked adapter and indexing primer products to ensure they produce sequencing libraries that meet our high-quality standards.
Our leadership in NGS adapter synthesis made us the clear choice when Illumina sought a partner to develop the next generation of indexed adapters to improve sample multiplexing. These adapters contain Illumina’s unique dual indexes (UDIs), which mitigate sample misassignment due to index hopping. We manufacture UDI adapters for Illumina and are also licensed to sell your own custom designs containing these UDI sequences.
For more information, or to order custom adapters, please contact us.
| Indexing Option | xGen UDI-UMI Adapters | xGen Stubby Adapter and UDI Primers | xGen Normalase UDI Primers | xGen UDI Primers | xGen Normalase CDI Primers | xGen CDI Primers | Custom NGS adapters |
|---|---|---|---|---|---|---|---|
| Configuration | UDI-UMI | UDI | UDI | UDI | CDI | CDI | Single index, combinatorial dual indexing, UDI, or UDI-UMI |
| Sample index length | 8 base, unique sample indexes | 8 base, unique sample indexes | 10 base, unique sample indexes | Both 8 and 10 base available, unique sample indexes | 8 base indexes | 8 base indexes | Configurable |
| Concentration | 15 µM | 15 µM (Stubby adapter), 10 µM ( each primer) | Proprietary | 20 µM total (10 µM each Primer)
| Proprietary | 12 µM total (6 µM each primer after arraying) | 15 µM (Adapter), 20 µM total (10 µM each Primer) |
| Shipping container | 96-well plates, single use | Tubes (Stubby Adapter), Single-use plates (Primers) | 96-well plates, single use | 96-well plates, single use UDI 10 nt Primers (2nm) plates are bulk/multi-use | Tubes for each i7 and i5 index | Tubes for each i7 and i5 index | Tubes, Matrix® plates†, or single-use plates |
| Reactions | 1 reaction per adapter | 1 reaction per primer pair | 1 rxn per primer pair | 1 rxn per primer pair for 96-well plates, single use | 1 rxn per primer pair | 1 rxn per primer pair | Varies |
| Illumina platform compatibility | 2- and 4- color systems | 2- and 4- color systems | 2- and 4- color systems | 2- and 4- color systems | 2- and 4- color systems | 2- and 4- color systems | Designs for any application |
Choose from a variety of options that personalize the adapters to your NGS workflow for Illumina sequencing. Order Custom NGS Adapters when you need to:
- Scale up your experiments—get the delivery format (volume, plate, single-use) and volume-pricing that best suit your NGS study.
- Expand your set of barcodes to increase sample throughput—augment limited sets of stocked adapters. Our design tool will support uploading up to 2000 custom index sequences!
- Supplement commercial library kit components—create your own custom library prep workflow that fits your experimental design.
The Custom Adapter Configurator tool helps you design the adapters best suited to your experiments by walking you through these choices:
- Library prep method
- Configuration (SI, CDI, UDI, UDI-UMI)
- Attachment style
- Indexing strategy
- Number of indexes
- Size (reaction number or nmol)
- Delivery format (tubes, plates)
- Methylated/unmethylated
- TruGradeTM processing
Table 2. Suggested barcode configuration based on NGS application.
| Application | Best choice | Alternatives |
|---|---|---|
| Germline characterization SNPs, indels, crude CNV; no concern of sample crosstalk | UDI | SI, CDI, or UDI-UMI |
| Germline characterization where sample crosstalk is important Translational research, inherited genetics, infectious disease research | UDI | UDI-UMI |
| Low-level mutation detection, down to ~1% frequency Cancer, mosaicism, other diseases where somatic mutations are important | UDI-UMI |
Resources
Frequently asked questions
What plate sealing guidelines should I follow when fewer than 16 or 96 libraries are prepared?
You should use an adhesive plate seal over the punctured but unused wells to prevent future cross‑contamination. Do not heat‑seal the plate twice.
What are the new names of IDT next generation sequencing products?
After IDT acquired Swift Biosciences, some of the IDT next generation sequencing products were renamed.
The following table lists the previous IDT product name corresponding with its new name, hyperlinked to the appropriate product page.
| Previous IDT product name | New IDT product name |
|---|---|
| xGen™ Prism™ DNA Library Prep Kit | xGen cfDNA & FFPE DNA Library Prep MC |
| COVID-19 Capture Bundle | xGen SARS-CoV-2 Hyb Panel Bundle |
| xGen CNV Backbone Panel—Tech Access | xGen CNV Backbone Hyb Panel |
| xGen Human ID Research Panel | xGen Human ID Hyb Panel |
| xGen Human mtDNA Research Panel v1.0 | xGen Human mtDNA Hyb Panel |
| xGen Exome Research Panel v2 | xGen Exome Hyb Panel v2 |
| Lotus™ DNA Library Prep Kit | xGen DNA Library Prep Kit MC or xGen DNA Library Prep Kit EZ |
| Custom or predesigned rhAmpSeq™ Library Prep Kits | Discontinued, but technology is available via rhAmpSeq for CRISPR Analysis System |
| Discovery Pools | xGen Custom Hyb Panels and xGen Custom Hyb Panels—Accel |
| Lockdown™ Probes Functional QC Discovery Pool | xGen Custom Hyb Panels—Production (available with and without functional testing) |
| xGen Predesigned Gene Capture Pools | xGen Custom Hyb Panels |
What are the new names of the Swift products?
The Swift products were rebranded and now belong to the xGen™ NGS product line. The following lists the old Swift product name, and then the new name hyperlinked to its current product page.
| Swift product name | New IDT product name |
|---|---|
| Accel-NGS® Adaptase® Module for Single-Cell Methyl-Seq | xGen Adaptase™ Module |
| Normalase® Amplicon Panels (SNAP) Core | xGen Amplicon Core |
| SARS-CoV-2 Additional Genome Coverage Panel | xGen SARS-CoV-2 Expanded Amplicon Panel |
| SARS-CoV-2 S Gene Panel | xGen SARS-CoV-2 SGene Amplicon Panel |
| SNAP Set 1A Combinatorial Dual Indexing Primers | xGen Amplicon CDI Primers |
| Swift Combinatorial Dual Indexing Primers | xGen CDI Primers |
| Swift Normalase® Combinatorial Dual Indexing Primers | xGen Normalase CDI Primers |
| Accel-NGS 1S Plus DNA Library | xGen ssDNA & Low-input DNA Library Prep |
| Swift 2S Sonic DNA Library | xGen DNA Library Prep MC |
| Swift 2S Sonic Flexible DNA Library | xGen DNA Library Prep MC UNI |
| Swift 2S Turbo v2 | xGen DNA Library Prep EZ |
| Swift 2S Turbo Flexible v2 DNA Library | xGen DNA Library Prep EZ UNI |
| Accel-NGS Methyl-Seq DNA Library | xGen Methyl-Seq Library Prep |
| Swift Normalase | xGen Normalase Module |
| Swift Rapid RNA Library | xGen RNA Library Prep Kit |
| Swift RNA Library | xGen Broad-Range RNA Library Prep |
Why are the xGen™ UDI Plate 2, 8 nt and the xGen CDI Primers a different concentration than other xGen Indexing Primers?
These two specific products were brought into the xGen portfolio from Swift Biosciences. To preserve the configuration of the products that support Swift's legacy customers, we did not alter the concentration or any other specifications, we only rebranded these products using new names.
Which sequencing platform should be used for the xGen™ Stubby Adapter–UDI Primers?
NGS libraries constructed using the stocked xGen Stubby Adapter–UDI Primers are Illumina TruSeq®-compatible and should be sequenced on an Illumina platform with our UDI-UMI Adapters and Normalase™ Indexing options.
Our Custom Adapter Configurator Tool helps you to create custom indexing options for not only Illumina TruSeq®- compatible workflows, but also Illumina Nextera®-compatible workflows.
TruSeq® and Nextera® are registered trademarks of Illumina, Inc., used with permission. All rights reserved.
Can I use the xGen™ UDI Plate 1, 8 nt and xGen UDI Plate 2, 8 nt together?
No, we do not recommend mixing those indexes. The xGen UDI Plate 1 is IDT's legacy UDI primer set, while the xGen UDI Plate 2 is Swift's legacy UDI set.
If you have specific questions on which samples can be multiplexed, please reach out to our Scientific Applications Support team for answers.
Can I use my own library amplification primers or indexing amplification primers for Normalase™ PCR?
No, Normalase PCR requires replacing your amplification primers to condition your libraries for the downstream enzymology. Reagent R5 replaces standard library amplification primers for those libraries indexed by adapter ligation.
Reagent R6 with xGen™ Normalase CDI primers, or Reagent R7 with Normalase UDI primers, replace your indexing primers for libraries indexed by PCR.
What amount of xGen™ UDI-UMI Adapters should I use for library prep?
The amount of xGen UDI-UMI Adapters you use depends on the library prep workflow and the amount of starting sample used to perform library prep. We recommend checking your specific protocol for the optimal amount of adapter to use.
For the xGen DNA Library Prep Kits, use 5 µL of xGen UDI-UMI Adapters (diluted according to the table below) for the ligation reaction.
Note: If diluting the xGen Stubby Adapter is required, we recommend using Nuclease-Free Duplex Buffer.
| DNA input (ng) | Adapter dilution | Stock concentration (µM) |
|---|---|---|
| ≥25 | No dilution | 15 |
| 10 | 10-fold (1:10) | 1.5 |
| 1 | 20-fold (1:20) | 0.75 |
Reach out to our Scientific Applications Support team if you are unsure.
Are methylated UDI-UMI adapters available?
We do not have them available as a stocked product, but they are able to be ordered through our Custom Adapter Configurator design tool.
Can adapter ligation be carried out in the provided 96-well plate of xGen™ UDI-UMI Adapters?
No, this is not recommended.
All our plate-based indexing products contain overage to allow for pipetting error, so we recommend pipetting directly out of the index plate and into a separate reaction plate, or tube.
Can xGen™ UDI-UMI Adapters be used for RNA-seq?
The xGen™ RNA Library Prep Kit includes a stubby adapter in the workflow so the stocked UDI-UMI adapters are not compatible with those kits.
For customers using homebrew, or other commercial RNA-seq workflows that utilize TA-ligation, these UDI-UMI adapters are compatible. However, it is important to note that stranded RNA-seq libraries constructed using xGen UDI-UMI Adapters have the insert strandedness flipped (i.e., to the opposite strand) when compared to standard Illumina TruSeq™
adapters. As such, this strand flip must be handled during data analysis.
Disclaimer: TruSeq™ and Nextera™ are registered trademarks of Illumina, Inc., used with permission. All rights reserved.
How do I sequence the UMI in xGen™ UDI-UMI Adapters?
The UMI (dark blue in the illustration below) is positioned next to the i7 index (light orange) so that it will be sequenced as part of the i7 index read.
So, to sequence the UMI, increase the number of cycles for the i7 index read by 9 cycles. Refer to the IDT Master Index List for guidelines concerning the xGen™ UDI-UMI adapters.

Do the xGen™ Stubby Adapter–UDI Primers contain UMIs?
No, the xGen Stubby Adapter–UDI Primers do not contain UMIs.
The structure of the final library created using the xGen Stubby Adapter–UDI Primers is diagrammed below:

What is the shelf life and recommended storage conditions for the xGen™ Stubby Adapter–UDI Primers?
Store all xGen™ indexing products at –20°C. The expiration date is on the kit label and the Certificate of Analysis, which can be found by searching the batch/lot number on the kit label.
What amount of xGen™ Stubby Adapter–UDI Primers should I use for library prep?
This can vary by library prep method; consult the specific protocol you are using for the exact input range.
Most of IDT’s library prep kits use a 5 μL input for either adapter ligation (full-length or stubby), or indexing PCR.
Are the xGen™ Stubby Adapter and Unique Dual Index Primer Pairs methylated?
The xGen™ Stubby Adapter is methylated.
However, the xGen Unique Dual Index Primer Pairs are not methylated. Bisulfite conversion can be carried out after ligation of the stubby adapter, but before library amplification.
Custom NGS Adapters can be purchased as 16 or 96 reactions. How many nanomoles are in 1 reaction?
For Illumina TruSeq™–Compatible Full-length Adapters, 1 reaction = 0.075 nmol.
For Illumina TruSeq™–Compatible Stubby Adapter, 1 reaction = 0.075 nmol (use with Illumina TruSeq™–Compatible Primers).
For Illumina TruSeq™–Compatible Primers, 1 reaction = 0.050 nmol (use with Illumina TruSeq™– Compatible Stubby Adapter).
For Illumina Nextera™–Compatible Indexing Primers, 1 reaction = 0.050 nmol.
TruSeq™ and Nextera™ are registered trademarks of Illumina, Inc., used with permission. All rights reserved.
(Visit the IDT Terms, usage, warranty & licensing page for specific information about Illumina trademarks)
What are the sequences for the IDT and Illumina® indexes: IDT 8, IDT 10, ILMN 8, ILMN 10?
These index sequences will be provided once your order has been placed.
You can also access the index sequence files for IDT 8/10 and ILMN 8 in the Custom Adapter Configurator tool. We are not permitted to share the ILMN 10 sequences pre-sale.
Contact our Scientific Applications Support team for further information.
TruSeq® and Nextera® are registered trademarks of Illumina, Inc., used with permission. All rights reserved.
GMP refers to products manufactured under ISO 13485: 2016 QMS. Purchaser is solely responsible for all decisions regarding the use of these products and any associated regulatory or legal obligations for their legal marketing.