Electrophoresis with agarose and polyacrylamide gels is one of the most widely used tools in molecular biology. Gels provide a simple, low-cost way to separate nucleic acids based on size for quantification and purification.
Agarose vs. polyacrylamide gels
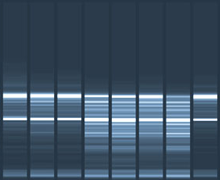
Agarose gels can be used to resolve large fragments of DNA. Polyacrylamide gels are used to separate shorter nucleic acids, generally in the range of 1−1000 base pairs, based on the concentration used (Figure 1). These gels can be run with or without a denaturant. Gels that are run without a denaturant are referred to as native gels. The DNA or RNA will migrate at different rates, depending on its secondary structure. Native gels allow the DNA or RNA to remain double stranded. Adding a denaturant to the gel, such as urea, will generally make all of the nucleic acids single stranded. Secondary structure will not form in denaturing gels and, therefore, only the length of the DNA will affect mobility.
Different concentrations of agarose and acrylamide are necessary to optimize resolution of nucleic acids with different lengths. Suggested concentrations are shown below in Table 1.
Table 1. Gel concentrations for size separation.
| Agarose gels | Polyacrylamide gels | ||
|---|---|---|---|
| % Agarose | Size range for optimum resolution (bp) | % Acrylamide | Size range for optimum resolution (bp) |
| 0.5 | 1,000−30,000 | 3.5 | 1,000−2,000 |
| 0.7 | 800−12,000 | 5 | 80−500 |
| 1.0 | 500−10,000 | 8 | 60−400 |
| 1.2 | 400−700 | 12 | 25−150 |
| 1.5 | 200−500 | 15 | 25−150 |
| − | − | 20 | 6−100 |
Safety considerations
Acrylamide is a potent neurotoxin and, in its powdered form, can easily be aerosolized. Make sure to wear the appropriate personal protection, including gloves and a mask, when weighing out the material. Many companies sell acrylamide dissolved in water or pre-cast gels. These products are slightly more costly but reduce the risk of acrylamide inhalation.
Ethidium bromide is the most common DNA stain available; it is also toxic if inhaled, decomposes when heated to produce toxic gases, and is suspected of causing genetic defects [1]. Always wear gloves and avoid microwaving liquids containing ethidium bromide. Non-mutagenic fluorescent dyes available as an alternative include Bio-Safe™ (Bio-Rad), SYBR-safe™ (ThermoFisher Scientific), and GelRed® Nucleic Acid Stain (Phenix Research Products). While more costly than ethidium bromide, these stains reduce the need for isolating and decontaminating gel electrophoresis stations. Note that ethidium bromide only intercalates into double-stranded DNA and, therefore, is not a good stain for single-stranded DNA analysis.
Tips for acrylamide gel electrophoresis
- Use fresh Ammonium Persulfate (APS)─APS catalyzes the polymerization of acrylamide. Using old APS or APS stored above -20°C will result in slow or incomplete polymerization. Keep small, fresh aliquots in the freezer.
- Know how your tracking dye(s) will migrate─In agarose gels, bromophenol blue and xylene cyanol will migrate at approximately 300 and 3000 bp, respectively. These dyes will migrate at different rates in acrylamide gels depending on the gel density. Table 2 provides the approximate migration rate in terms of the relative size of single-stranded/denatured DNA.
- TAE or TBE? Agarose gels commonly use Tris-Acetic Acid-EDTA (TAE) or Tris-Boric Acid-EDTA (TBE) buffers. TAE buffer has the advantage that it can be made in 50X stock solutions. However, it buffers less efficiently and, in some cases, its use results in smeared bands. TBE buffered gels yield sharper bands, particularly when using small-sized DNA fragments, and can be run at higher voltages. However, the borate in TBE can inhibit some enzymes—including T4 DNA ligase—in DNA purified from these gels.
- What voltage to use? Agarose gels can be run at a large range of voltages—from 0.25–7 V/cm. High voltages save time but can result in overheating of the gel, even leading to melting of low percentage agarose gels. High voltages can also cause band smearing, particularly of fragments >10 kb. The sharpest bands can be obtained by running gels in TBE overnight at 0.25–0.5 V/cm.
Table 2. Dye migration rates in acrylamide gels.
140| Nondenaturing gels | Denaturing gels | |||
|---|---|---|---|---|
| % Acrylamide | Bromophenol blue (nucleotides) | Xylene cyanol (nucleotides) | Bromophenol blue (nucleotides) | Xylene cyanol (nucleotides) |
| 3.5 | 100 | 460 | 55 | 210 |
| 5.0 | 65 | 260 | 35 | |
| 8.0 | 45 | 160 | 19 | 75 |
| 12.0 | 20 | 70 | 10 | 70 |
| 15.0 | 15 | 60 | 8 | 28 |
*Adapted from Sambrook J, Fritsch ER, Maniatis T. Molecular Cloning: A Laboratory Manual. 2nd ed. Cold Spring Harbor laboratory Press; 1989.
Troubleshooting gel electrophoresis
- Blurry bands? Too much DNA or excess salt will create smeared bands and/or streaking in the gel. Loading the correct amount of DNA (usually a maximum of 100−250 ng/mm well width) and desalting samples with a spin column prior to loading will prevent this.
- Bands in the wrong place? Do not heat nucleic acids before running on a native gel, and do not exceed 20 V/cm (measured from anode to cathode, rather than entire gel length) or allow the gel to exceed 30°C. For the sharpest bands, run the gel slowly, at 5 V/cm.
- Loading buffer floats away? Rinsing wells with running buffer just before loading is essential; failure to do so may prevent the loading mixture from sinking to the bottom of the well, resulting in an uneven band and delayed migration.


 Processing
Processing