| Alt-R S.p. Cas9 Nuclease | Alt-R S.p. HiFi Cas9 Nuclease | Alt-R S.p. Cas9 D10A Nickase | Alt-R S.p. Cas9 H840A Nickase | |
|---|---|---|---|---|
 |
 |
 |
 |
|
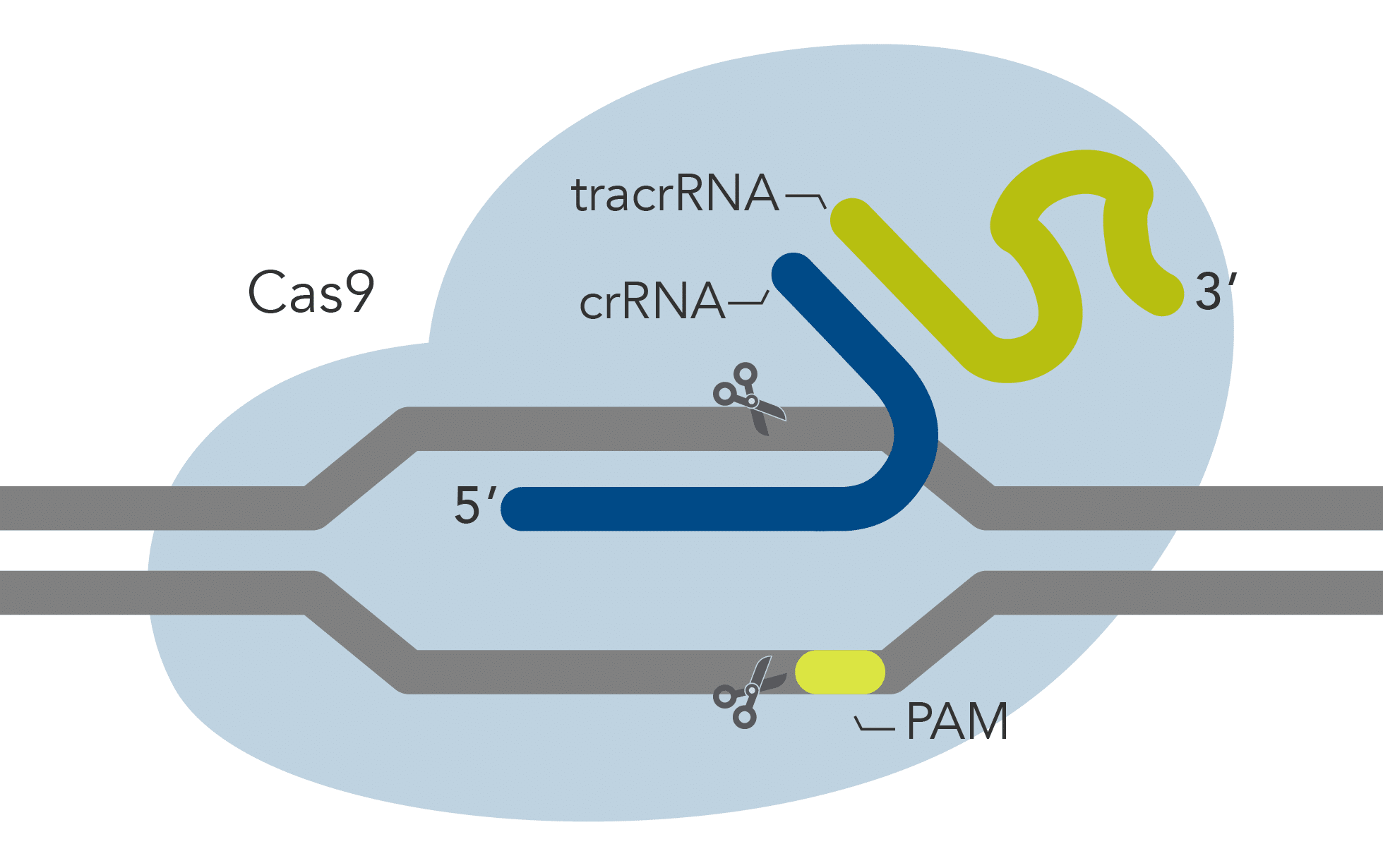
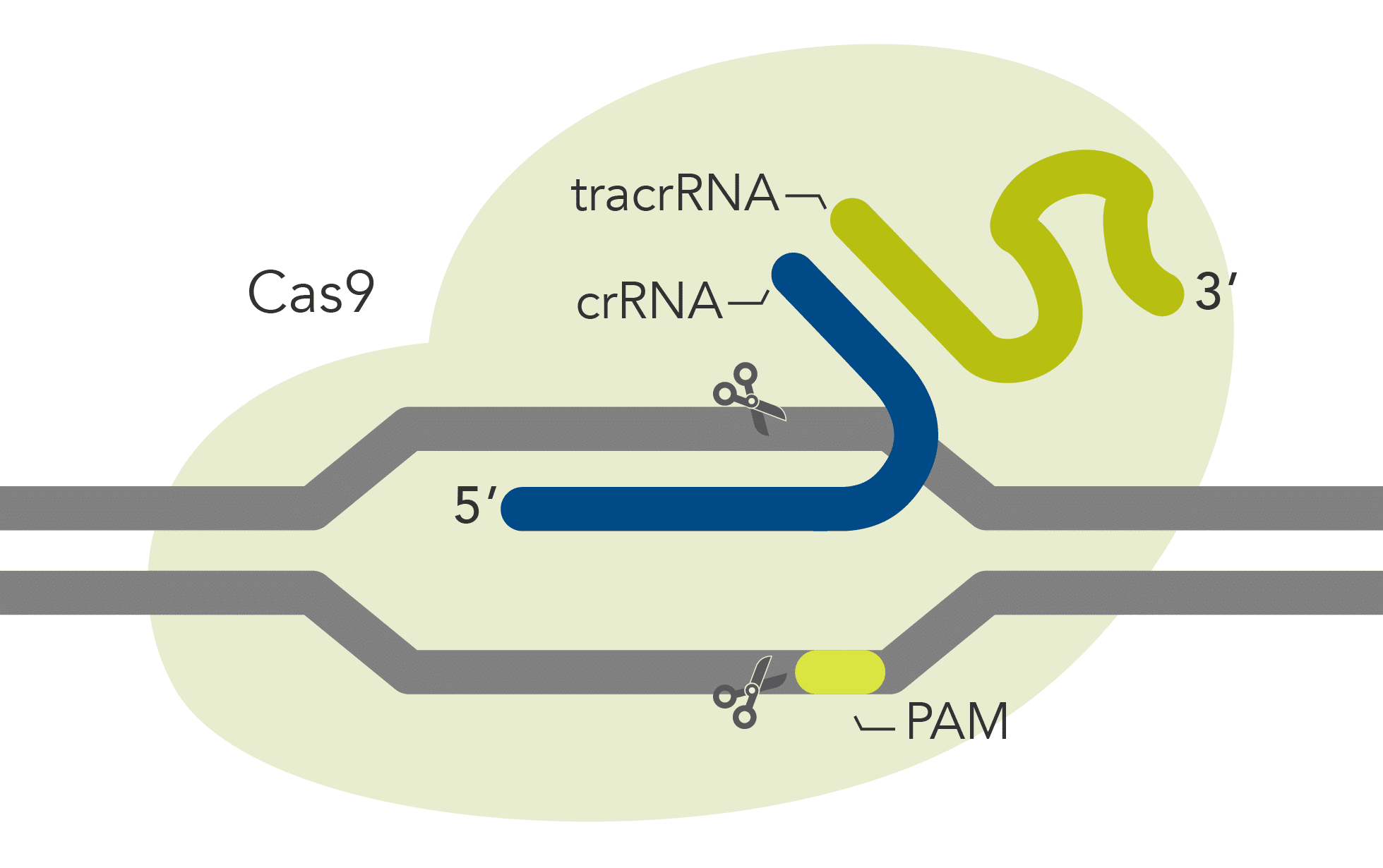
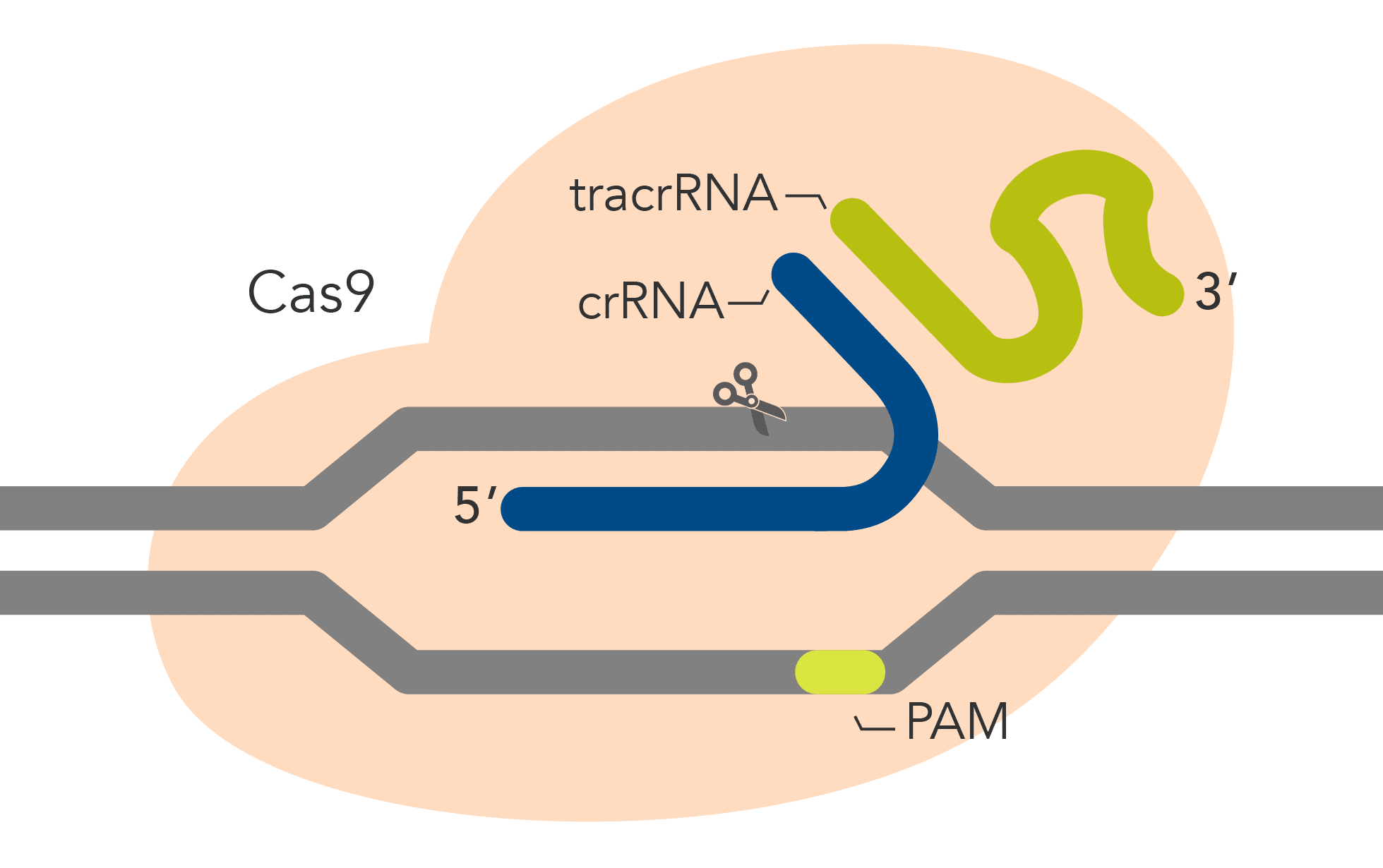
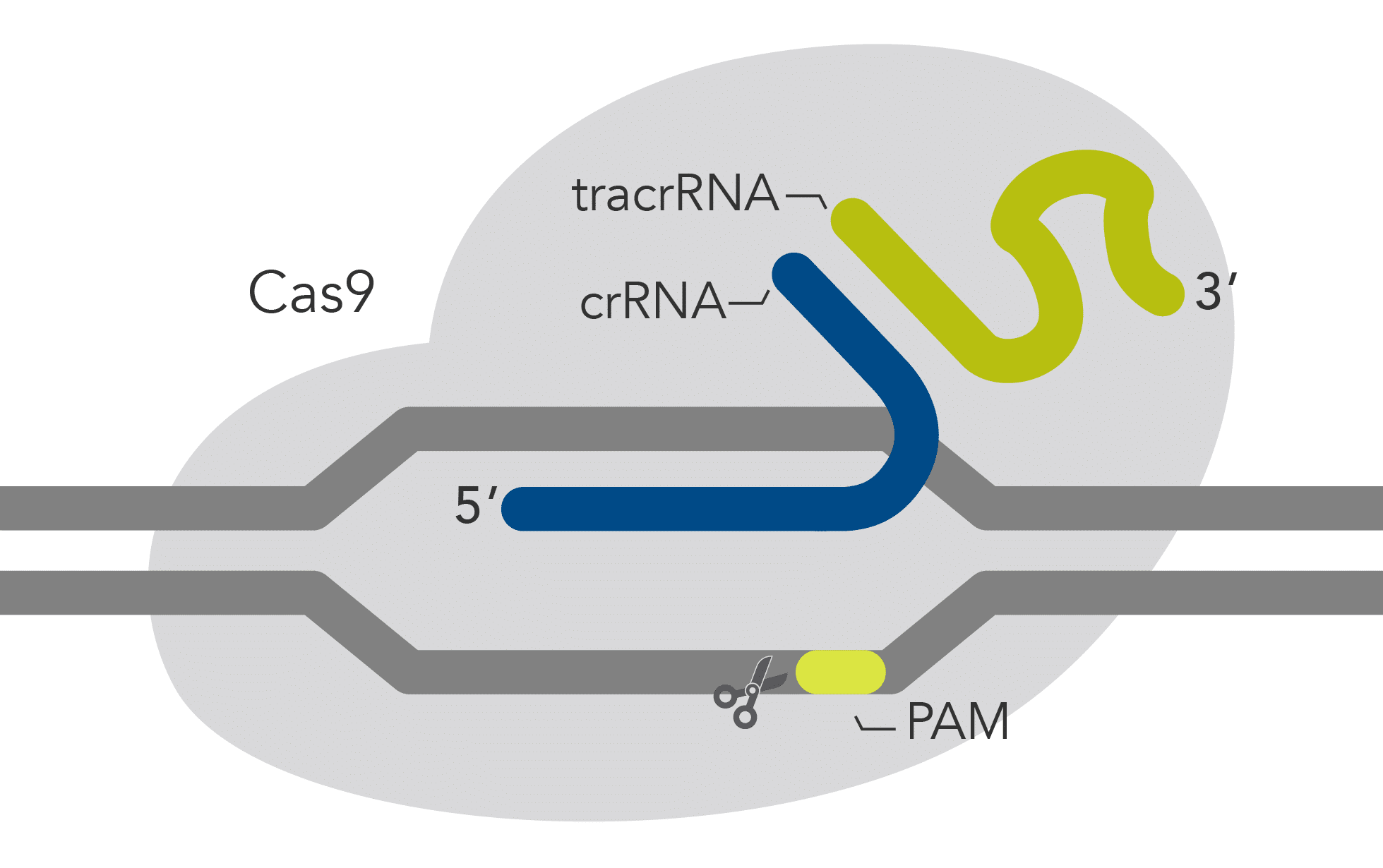
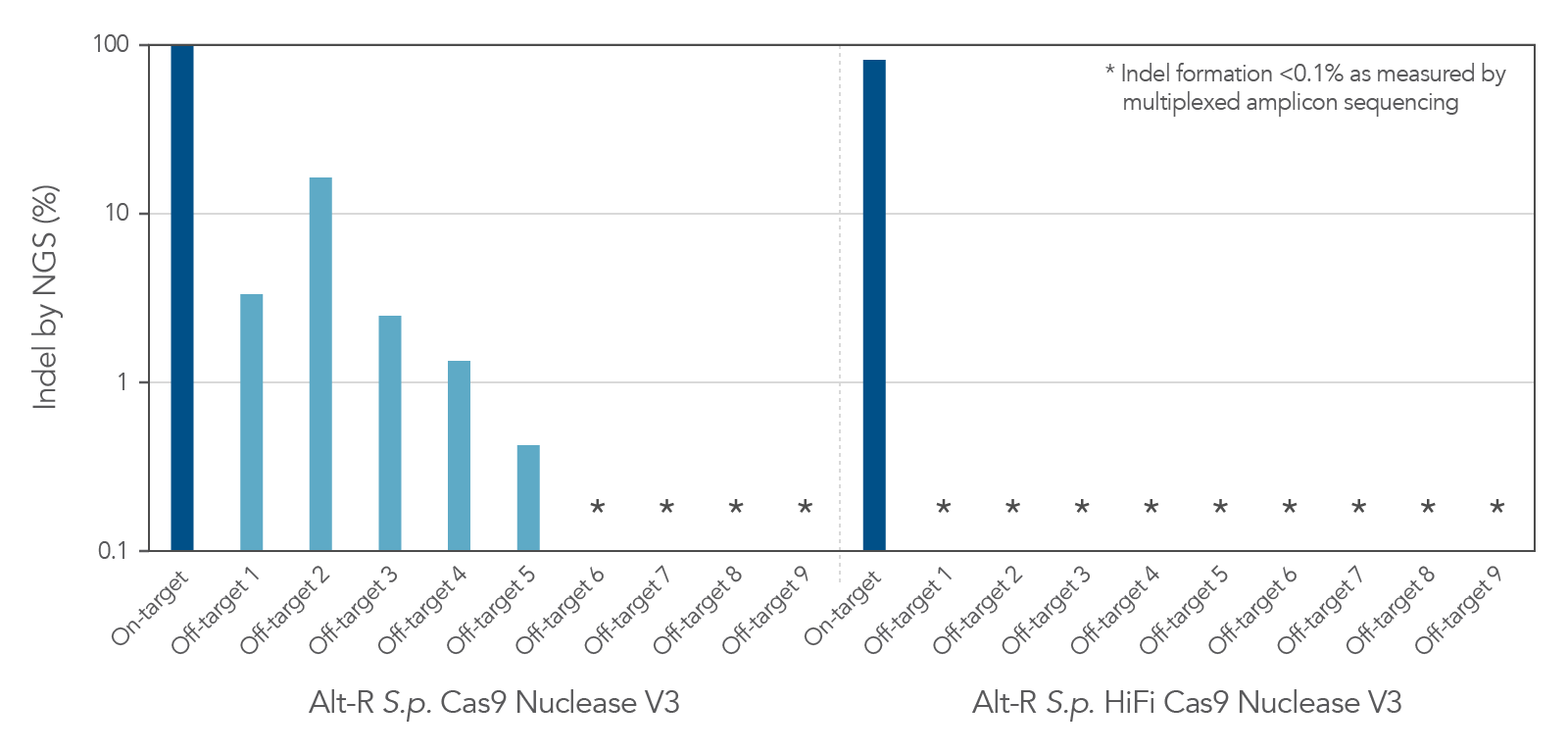
| Description | Wild-type Cas9 with high genome editing potency that is simple to use and economical | Cas9 variant with improved specificity based on reduced off-target effects, while preserving high on-target activity | Cas9 variant with a mutation in the RuvC domain that disables cleavage of the non-target strand | Cas9 variant with a mutation in the HNH domain that disables cleavage of the target strand |
| DNA cleavage | Both strands | Both strands | Target strand | Non-target strand |
| Suggested use | First choice for most CRISPR genome editing projects | Ideal for experiments sensitive to off-target events | May be beneficial for homology-directed repair (HDR) experiments, but requires two suitable cutting sites within an optimal distance | |
| Molecular weight | 162,200 g/mol | |||
| Amount provided | 100 μg or 500 μg | |||
| Concentration | 10 mg/mL (62 μM) in 50% glycerol | |||
| Shipping conditions | Dry ice | |||
| Storage conditions | -20°C at stock concentration | |||
| Dilution | Dilute in a medium (such as OptiMEM®) or Phosphate-buffered saline (PBS) before use | |||
Unless otherwise expressly indicated in a Certificate of Analysis, label or other documentation accompanying the Products, the Products are intended for research use only and subject in each and every case to the condition that such sale does not convey any license, expressly or by implication, to manufacture, duplicate or otherwise copy or reproduce any of the Products.
SpyFi is the property of Aldevron and all information and data presented are solely those of Aldevron. Unless agreed to in writing, IDT does not warrant the fitness or suitability of these products. Purchaser is solely responsible for all decisions regarding the use of these products and any associated legal or regulatory obligations.
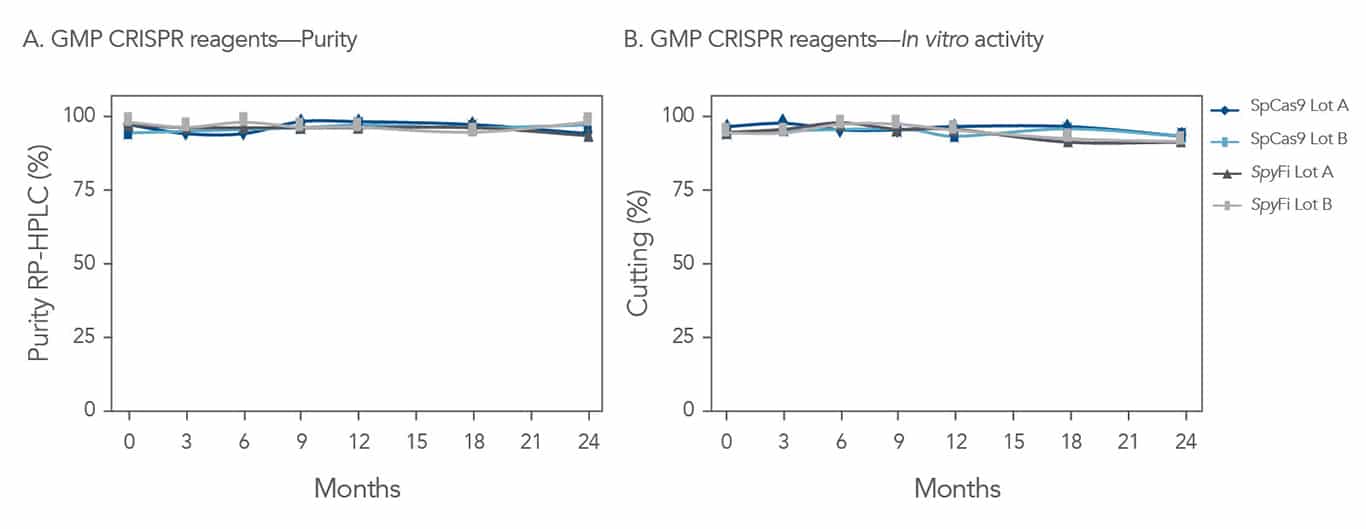
CGMP gRNA products are manufactured in accordance with ICH Q7, 21CFR210, 211, and parts of 600. IDT engineering runs and CGMP gRNA are for development and investigational use only. The performance characteristics of this product have not been established. This product is not intended to be used as final drug product. The purchaser is solely responsible for all decisions regarding the intended use of the product and any associated legal or regulatory obligations.
RUO24-3157_001